 The function returns a numpy memoryview, which works well enough in most cases. 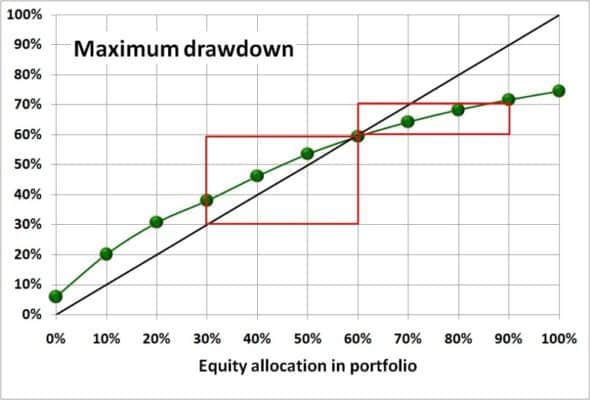
The function to call is cy_rolling_dd_custom_mv where the first argument ( ser) should be a 1-d numpy array and the second argument ( window) should be a positive integer. For typical use cases, the speedup vs regular python was ~100x or ~150x. There was a bit of work to do to make sure I'd properly typed everything (sorry, new to c-type languages). To handle NA's, you could preprocess the Series using the fillna method before passing the array to rolling_max_dd.įor the sake of posterity and for completeness, here's what I wound up with in Cython. For example, with window_length = 200, it is almost 13 times faster. The speedup is better for smaller window lengths. Rolling_max_dd is about 6.5 times faster. 
In : %timeit my_rmdd = rolling_max_dd(s.values, window_length, min_periods=1) Timing comparison, with n = 10000 and window_length = 500: In : %timeit rolling_dd = pd.rolling_apply(s, window_length, max_dd, min_periods=0) The green dots are computed by rolling_max_dd. The plot shows the curves generated by your code. My_rmdd = rolling_max_dd(s.values, window_length, min_periods=1) Rolling_dd = pd.rolling_apply(s, window_length, max_dd, min_periods=0)ĭf.columns = Running_max_y = np.maximum.accumulate(y, axis=1) Pad = np.empty(window_size - min_periods) Returns an 1d array with length `len(x) - min_periods + 1`. `min_periods` should satisfy `1 <= min_periods <= window_size`. Y = as_strided(x, shape=(x.size - window_size + 1, window_size),ĭef rolling_max_dd(x, window_size, min_periods=1): `min_periods` should satisfy `1 > x = np.array() """Compute the rolling maximum drawdown of `x`. def rolling_max_dd(x, window_size, min_periods=1): Once we have this windowed view, the calculation is basically the same as your max_dd, but written for a numpy array, and applied along the second axis (i.e. windowed_view is a wrapper of a one-line function that uses _tricks.as_strided to make a memory efficient 2d windowed view of the 1d array (full code below). Here's a numpy version of the rolling maximum drawdown function. I think that could be a very fast solution if implemented in Cython. If anyone is interested, the "bespoke" algorithm I alluded to in my post is rolling_dd_custom. It shows how some of the approaches to this problem relate, checks that they give the same results, and shows their runtimes on data of various sizes. Or, perhaps, that someone might want to have a look at my "handmade" code and be willing to help me convert it to Cython.įor anyone who wants a review of all the functions mentioned here (and some others!) have a look at the iPython notebook at: I was hoping someone had tried this before. But I'm not currently fluent enough in Cython to really know how to begin attacking this from that angle. 
I think it's because of all the looping overhead in Python/Numpy/Pandas. It does save some time, but not a whole lot, and not nearly as much as should be possible. Is there a particularly slick algorithm in pandas or another toolkit to do this fast? I took a shot at writing something bespoke: it keeps track of all sorts of intermediate data (locations of observed maxima, locations of previously found drawdowns) to cut down on lots of redundant calculations. It works like so: rolling_dd = pd.rolling_apply(s, 10, max_dd, min_periods=0) This is easy to do using pd.rolling_apply. for each step, I want to compute the maximum drawdown from the preceding sub series of a specified length. Now say I'm interested in computing the rolling drawdown of this Series. S = pd.Series(np.random.randn(n).cumsum())Īs expected, max_dd(s) winds up showing something right around -17.6. Let's set up a brief series to play with to try it out: np.ed(0) It takes a small bit of thinking to write it in O(n) time instead of O(n^2) time. It's pretty easy to write a function that computes the maximum drawdown of a time series. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
